10x merge targets on human/mouse datasets
Viktor Petukhov
2018-01-23
Source file: notebooks/cell_barcode_correction/merge_targets_mixture.Rmd
Last updated: 2018-05-31
Code version: 35d0d5a
library(ggplot2)
library(ggrastr)
library(dplyr)
library(parallel)
library(dropestr)
library(dropEstAnalysis)
library(Matrix)
theme_set(theme_base)Load data
kDataPath <- '../../data/dropest/'
kTablesPath <- '../../output/tables/'
k10xSubfolders <- c(poisson='est_01_14_precise/', real='est_01_14_barcodes/',
unmerged='est_01_14_unmerged/', merge_all='est_01_16_merge_all/')
kDropSeqSubolders <- c(poisson='est_01_16_precise/', unmerged='est_01_16_unmerged/',
merge_all='est_01_16_merge_all/')
kDataFiles <- list(
`10x`=paste0(kDataPath, '10x/hgmm_6k/', k10xSubfolders, "hgmm6k.rds") %>%
setNames(names(k10xSubfolders)),
drop_seq=paste0(kDataPath, 'dropseq/thousand/', kDropSeqSubolders, "thousand.rds") %>%
setNames(names(kDropSeqSubolders))
)holders <- mclapply(kDataFiles, function(paths) mclapply(paths, readRDS, mc.cores=4),
mc.cores=2)
validation_data <- mclapply(holders, function(hs) list(
merge_targets = lapply(hs, function(holder) unlist(holder$merge_targets)),
cms_raw = lapply(hs, `[[`, 'cm_raw'),
cms = lapply(hs, `[[`, 'cm')
), mc.cores=8)
validation_data$`10x`$cms_raw <- lapply(validation_data$`10x`$cms_raw,
function(cm) cm[grep("^[^;]+$", rownames(cm)),])
validation_data$`10x`$cms <- lapply(validation_data$`10x`$cms,
function(cm) cm[grep("^[^;]+$", rownames(cm)),])
rm(holders)
invisible(gc())
# saveRDS(validation_data, paste0(kDataPath, 'human_mouse_mixture_validation_data.rds'))
# validation_data <- readRDS(paste0(kDataPath, 'human_mouse_mixture_validation_data.rds'))10x
umis_per_cb <- Matrix::colSums(validation_data$`10x`$cms$unmerged) %>% sort(decreasing=T)
real_cbs <- names(umis_per_cb)[1:6000]
PlotCellsNumberLine(umis_per_cb[1:10000])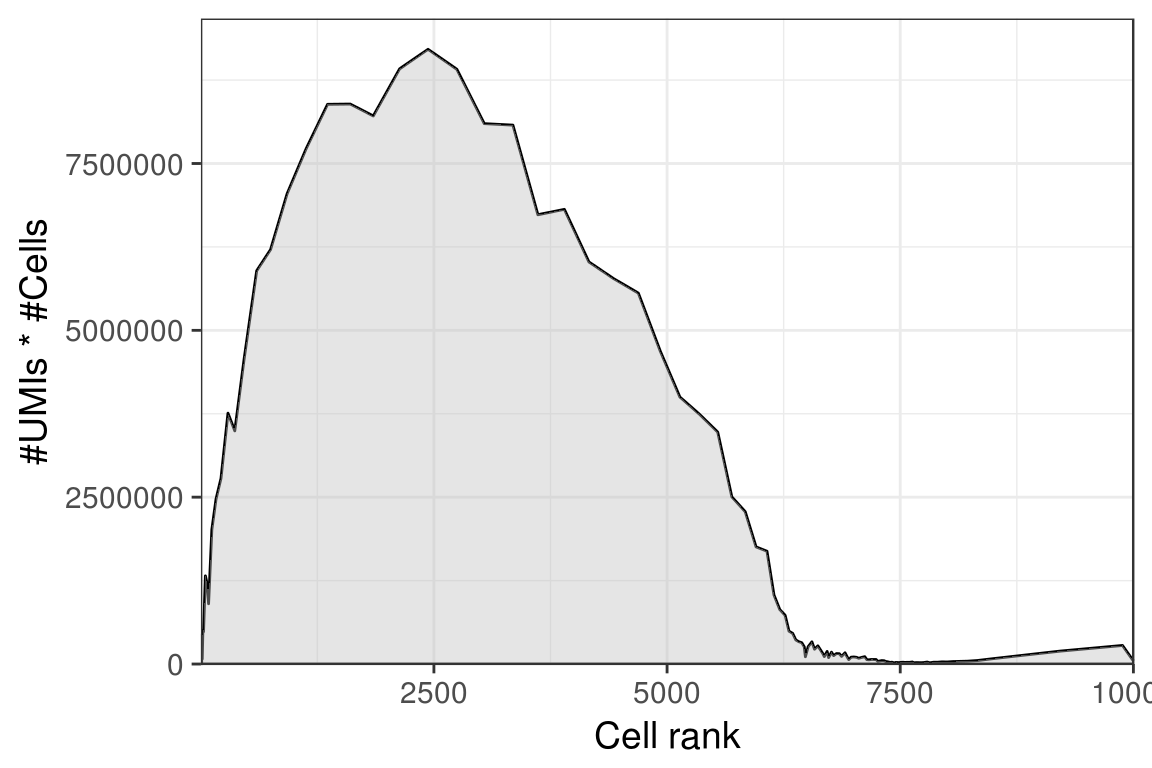
GeneSpeciesFromMixture <- function(cm, org1.marker, org1.name, org2.name) {
res <- ifelse(substr(rownames(cm), 1, nchar(org1.marker)) == org1.marker, org1.name, org2.name)
return(as.factor(res))
}
CellSpeciesFromMixture <- function(gene.species, cm) {
res <- levels(gene.species) %>%
lapply(function(l) cm[gene.species == l,] %>% Matrix::colSums())
res <- levels(gene.species)[as.integer(res[[1]] < res[[2]]) + 1] %>%
setNames(colnames(cm)) %>% as.factor()
return(res)
}gene_species <- GeneSpeciesFromMixture(validation_data$`10x`$cms_raw$unmerged,
'hg', 'Human', 'Mouse')
cell_species <- CellSpeciesFromMixture(gene_species,
validation_data$`10x`$cms_raw$unmerged)
table(cell_species[real_cbs])
Human Mouse
3260 2740 table(cell_species) / sum(table(cell_species))cell_species
Human Mouse
0.7844979 0.2155021 merge_targets <- lapply(validation_data$`10x`$merge_targets,
function(mt) mt[mt %in% real_cbs])
comparison_10x <- MergeComparisonSummary(merge_targets, cell_species, dataset="10x hgmm6k")
comparison_10x$`Merge type` <- c('Poisson', 'Known barcodes', 'Simple')
comparison_10x| Dataset | Merge type | #Merges | Fraction of mixed merges | Similarity to merge with barcodes |
|---|---|---|---|---|
| 10x hgmm6k | Poisson | 8999 | 0.58% | 99.74% |
| 10x hgmm6k | Known barcodes | 8985 | 0.62% | 100% |
| 10x hgmm6k | Simple | 21827 | 32.96% | 20.67% |
Drop-seq
umis_per_cb <- Matrix::colSums(validation_data$drop_seq$cms$unmerged) %>%
sort(decreasing=T)
real_cbs <- names(umis_per_cb)[1:1000]
PlotCellsNumberLine(umis_per_cb[1:5000])
gene_species <- GeneSpeciesFromMixture(validation_data$drop_seq$cms_raw$unmerged,
'HUMAN', 'Human', 'Mouse')
cell_species <- CellSpeciesFromMixture(gene_species,
validation_data$drop_seq$cms_raw$unmerged)
table(cell_species[real_cbs])
Human Mouse
586 414 table(cell_species)cell_species
Human Mouse
52885 9670 merge_targets <- lapply(validation_data$drop_seq$merge_targets,
function(mt) mt[mt %in% real_cbs])
comparison_drop_seq <- MergeComparisonSummary(merge_targets, cell_species,
dataset='Drop-seq thousand')
comparison_drop_seq$`Merge type` <- c('Poisson', 'Simple')
comparison_drop_seq| Dataset | Merge type | #Merges | Fraction of mixed merges | Similarity to merge with barcodes |
|---|---|---|---|---|
| Drop-seq thousand | Poisson | 15186 | 0.83% | NA% |
| Drop-seq thousand | Simple | 26154 | 8.74% | NA% |
complete_table <- rbind(comparison_10x, comparison_drop_seq)
write.csv(complete_table, paste0(kTablesPath, 'merge_comparison.csv'), row.names=F)Session information
| value | |
|---|---|
| version | R version 3.4.1 (2017-06-30) |
| os | Ubuntu 14.04.5 LTS |
| system | x86_64, linux-gnu |
| ui | X11 |
| language | (EN) |
| collate | en_US.UTF-8 |
| tz | America/New_York |
| date | 2018-05-31 |
| package | loadedversion | date | source | |
|---|---|---|---|---|
| 1 | assertthat | 0.2.0 | 2017-04-11 | CRAN (R 3.4.1) |
| 2 | backports | 1.1.2 | 2017-12-13 | CRAN (R 3.4.1) |
| 4 | bindr | 0.1 | 2016-11-13 | CRAN (R 3.4.1) |
| 5 | bindrcpp | 0.2 | 2017-06-17 | CRAN (R 3.4.1) |
| 6 | clisymbols | 1.2.0 | 2017-05-21 | CRAN (R 3.4.1) |
| 7 | colorspace | 1.3-2 | 2016-12-14 | CRAN (R 3.4.1) |
| 10 | digest | 0.6.15 | 2018-01-28 | cran (@0.6.15) |
| 11 | dplyr | 0.7.4 | 2017-09-28 | CRAN (R 3.4.1) |
| 12 | dropEstAnalysis | 0.6.0 | 2018-05-16 | local (VPetukhov/dropEstAnalysis@NA) |
| 13 | dropestr | 0.7.7 | 2018-03-17 | local (@0.7.7) |
| 14 | evaluate | 0.10.1 | 2017-06-24 | CRAN (R 3.4.1) |
| 15 | ggplot2 | 2.2.1 | 2016-12-30 | CRAN (R 3.4.1) |
| 16 | ggrastr | 0.1.5 | 2017-12-28 | Github (VPetukhov/ggrastr@cc56b45) |
| 17 | git2r | 0.21.0 | 2018-01-04 | cran (@0.21.0) |
| 18 | glue | 1.2.0 | 2017-10-29 | CRAN (R 3.4.1) |
| 22 | gtable | 0.2.0 | 2016-02-26 | CRAN (R 3.4.1) |
| 23 | highr | 0.6 | 2016-05-09 | CRAN (R 3.4.1) |
| 24 | htmltools | 0.3.6 | 2017-04-28 | CRAN (R 3.4.1) |
| 25 | knitr | 1.20 | 2018-02-20 | cran (@1.20) |
| 26 | labeling | 0.3 | 2014-08-23 | CRAN (R 3.4.1) |
| 27 | lattice | 0.20-35 | 2017-03-25 | CRAN (R 3.4.1) |
| 28 | lazyeval | 0.2.1 | 2017-10-29 | CRAN (R 3.4.1) |
| 29 | magrittr | 1.5 | 2014-11-22 | CRAN (R 3.4.1) |
| 30 | Matrix | 1.2-12 | 2017-11-16 | CRAN (R 3.4.1) |
| 32 | munsell | 0.4.3 | 2016-02-13 | CRAN (R 3.4.1) |
| 34 | pkgconfig | 2.0.1 | 2017-03-21 | CRAN (R 3.4.1) |
| 35 | plyr | 1.8.4 | 2016-06-08 | CRAN (R 3.4.1) |
| 36 | R6 | 2.2.2 | 2017-06-17 | CRAN (R 3.4.1) |
| 37 | Rcpp | 0.12.17 | 2018-05-18 | cran (@0.12.17) |
| 38 | rlang | 0.1.4 | 2017-11-05 | CRAN (R 3.4.1) |
| 39 | rmarkdown | 1.9 | 2018-03-01 | CRAN (R 3.4.1) |
| 40 | rprojroot | 1.3-2 | 2018-01-03 | cran (@1.3-2) |
| 41 | scales | 0.5.0 | 2017-08-24 | CRAN (R 3.4.1) |
| 42 | sessioninfo | 1.0.0 | 2017-06-21 | CRAN (R 3.4.1) |
| 44 | stringi | 1.1.7 | 2018-03-12 | cran (@1.1.7) |
| 45 | stringr | 1.3.0 | 2018-02-19 | cran (@1.3.0) |
| 46 | tibble | 1.3.4 | 2017-08-22 | CRAN (R 3.4.1) |
| 49 | withr | 2.1.2 | 2018-03-15 | cran (@2.1.2) |
| 50 | yaml | 2.1.18 | 2018-03-08 | cran (@2.1.18) |
This R Markdown site was created with workflowr